Antibodies show evidence of significant epistasis in biophysical properties but not binding
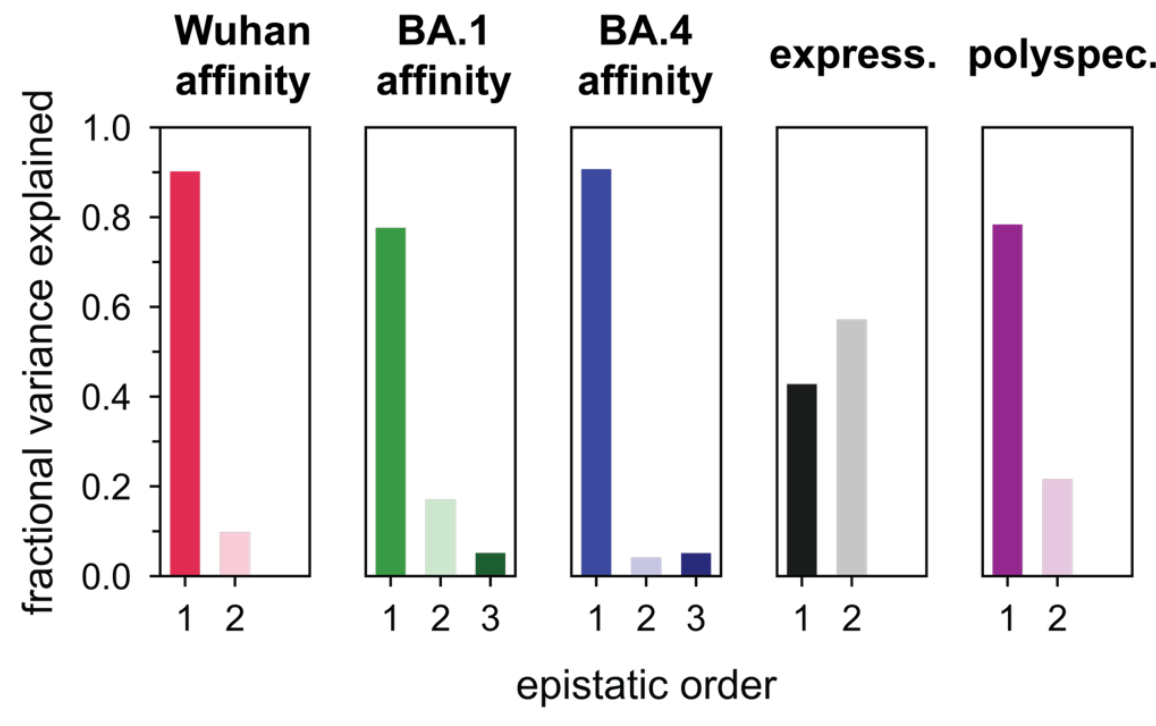
Antibodies undergo somatic mutation during affinity maturation in ways that show evidence of epistasis in biophysical properties like expression and polyspecificity, but not binding [1][2]. In a study by Kirby et al 2025[1:1] on 12 anti-SARS-CoV-2 antibodies using deep mutational scanning and yeast display, no combination of two point mutations showed greater than 30% difference in binding affinity compared to the sum of two individual mutations in isolation. Likewise, Tharp et al 2026 saw linear models explain 80-90% of single-antigen binding in their data. However, the latter saw that second-order models were necessary to model expression and polyspecificity.
References
Kirby, M. B., Petersen, B. M., Faris, J. G., Kells, S. P., Sprenger, K. G., & Whitehead, T. A. (2025). Retrospective SARS-CoV-2 human antibody development trajectories are largely sparse and permissive. Proceedings of the National Academy of Sciences, 122(4). https://doi.org/10.1073/pnas.2412787122 ↩︎ ↩︎
Tharp, C. R., Catalano, C., Khalifeh, A., Ghaffari-Kashani, S., Huang, R., Kang, G., Scapin, G., & Phillips, A. M. (2026). Biophysical trade-offs in antibody evolution are resolved by conformation-mediated epistasis. openRxiv. https://doi.org/10.64898/2026.03.12.711465 ↩︎