Antibody-antigen complex prediction by AF3-generation methods is data-limited
The recent report of Protenix-v1[1] contained an architecturally identical second model trained with a more recent train:test split. Unlike other outstanding problems in biomolecular structure prediction, antibody-antigen complex prediction showed remarkable improvement in the fraction of models achieving a cutoff of DockQ≥0.23, shown in the third column, bottom row:
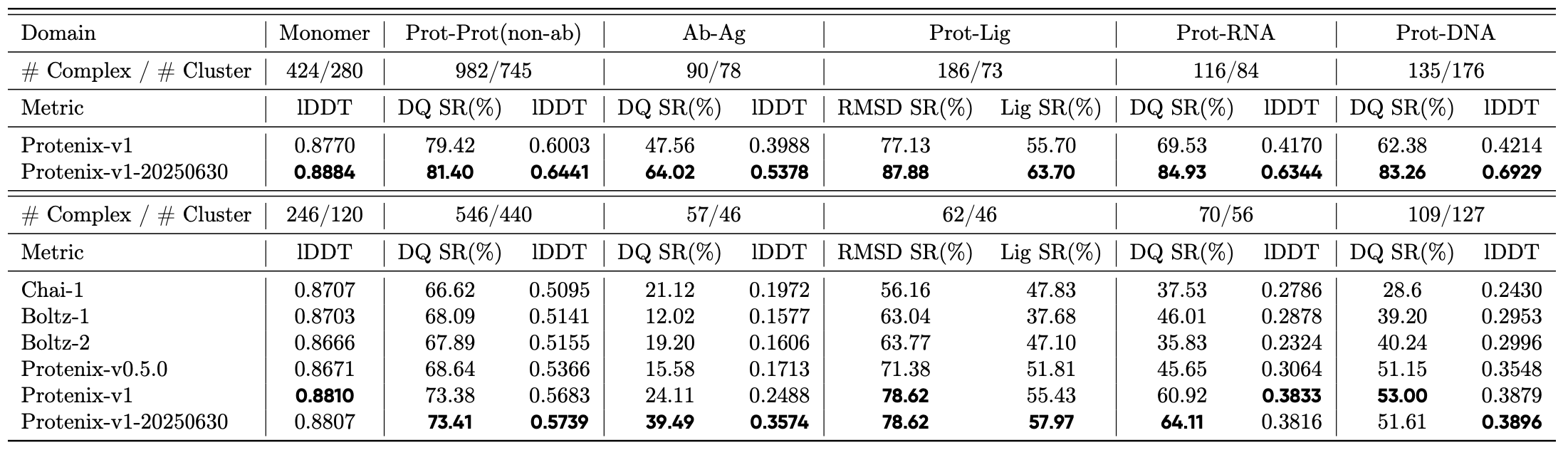
In contrast, other types of molecules, such as small molecules and nucleic acids, showed no improvements despite several years of additional data, suggesting fundamental architectural improvements are needed. This result was also implied by the IsoDDE white paper[2], which presumably does have such changes.
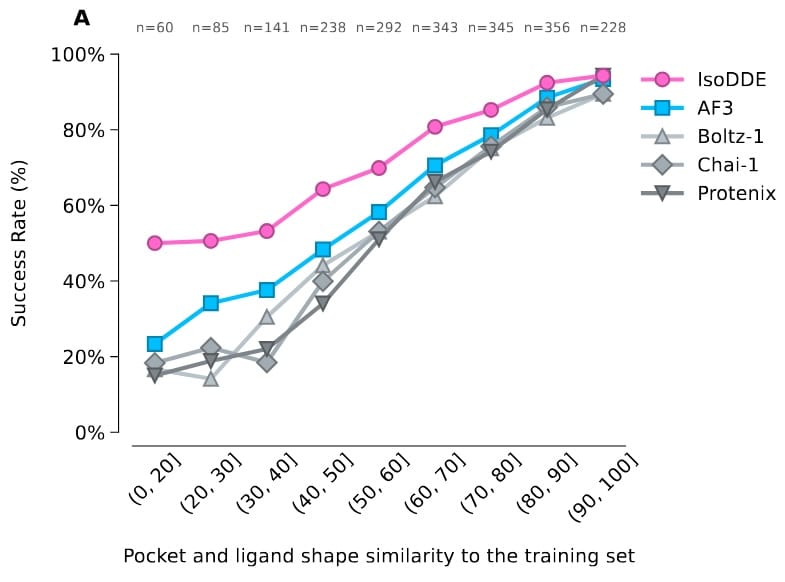
References
Zhang, Y., Gong, C., Zhang, H., Ma, W., Liu, Z., Chen, X., Guan, J., Wang, L., Yang, Y., Xia, Y., & Xiao, W. (2026). Protenix-v1: Toward High-Accuracy Open-Source Biomolecular Structure Prediction. openRxiv. https://doi.org/10.64898/2026.02.05.703733 ↩︎
Isomorphic Labs Team (2026). Accurate Predictions of Novel Biomolecular Interactions with IsoDDE. https://storage.googleapis.com/isomorphiclabs-website-public-artifacts/isodde_technical_report.pdf ↩︎