Conformational entropy could still matter in miniprotein binder design
Antibody V-regions improve their affinity for targets by both creating more high-energy interactions and reducing the conformational entropy of their antigen-binding loops[1][2]. Entropy's importance in antibody-antigen affinity seems obvious, given that loop residues largely mediate binding[3][4]. But for de novo-designed miniprotein binders, which often adopt helical bundle topologies and can now be created with high-enough success rates[5], the relevance of conformational entropy to overall design success remains unclear. This is because the inverse folding models used to design sequences onto backbones tend to excel at such highly structured backbones[6]. That said, a report from last year[7] found that directed evolution on de novo-designed miniprotein binders yielded affinity-enhancing mutations far from the interface, consistent with conformational stabilization; the authors attribute these mutations' effects to optimization of core packing.
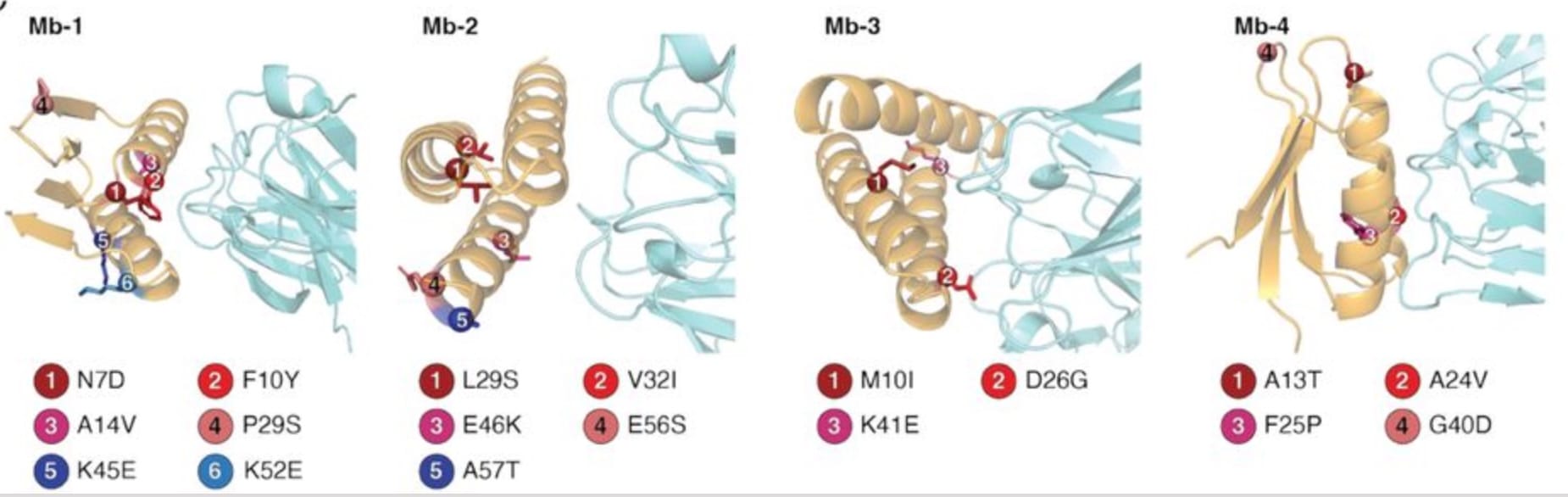
This suggests that, to some degree, entropy and conformational rigidity continue to be something worth thinking about in the context of miniprotein binder design.
References
Fernández-Quintero, M. L., Loeffler, J. R., Bacher, L. M., Waibl, F., Seidler, C. A., & Liedl, K. R. (2020). Local and Global Rigidification Upon Antibody Affinity Maturation. Frontiers in Molecular Biosciences, 7. https://doi.org/10.3389/fmolb.2020.00182 ↩︎
Guloglu, B., & Deane, C. M. (2023). Specific attributes of the VL domain influence both the structure and structural variability of CDR-H3 through steric effects. Frontiers in Immunology, 14. https://doi.org/10.3389/fimmu.2023.1223802 ↩︎
Mikolajek, H., Weckener, M., Brotzakis, Z. F., Huo, J., Dalietou, E. V., Le Bas, A., Sormanni, P., Harrison, P. J., Ward, P. N., Truong, S., Moynie, L., Clare, D. K., Dumoux, M., Dormon, J., Norman, C., Hussain, N., Vogirala, V., Owens, R. J., Vendruscolo, M., & Naismith, J. H. (2022). Correlation between the binding affinity and the conformational entropy of nanobody SARS-CoV-2 spike protein complexes. Proceedings of the National Academy of Sciences, 119(31). https://doi.org/10.1073/pnas.2205412119 ↩︎
Thorpe, I. F., & Brooks, C. L., III. (2007). Molecular evolution of affinity and flexibility in the immune system. Proceedings of the National Academy of Sciences, 104(21), 8821–8826. https://doi.org/10.1073/pnas.0610064104 ↩︎
Pacesa, M., Nickel, L., Schellhaas, C., Schmidt, J., Pyatova, E., Kissling, L., Barendse, P., Choudhury, J., Kapoor, S., Alcaraz-Serna, A., Cho, Y., Ghamary, K. H., Vinué, L., Yachnin, B. J., Wollacott, A. M., Buckley, S., Westphal, A. H., Lindhoud, S., Georgeon, S., … Correia, B. E. (2025). One-shot design of functional protein binders with BindCraft. Nature, 646(8084), 483–492. https://doi.org/10.1038/s41586-025-09429-6 ↩︎
Dauparas, J., Anishchenko, I., Bennett, N., Bai, H., Ragotte, R. J., Milles, L. F., Wicky, B. I. M., Courbet, A., de Haas, R. J., Bethel, N., Leung, P. J. Y., Huddy, T. F., Pellock, S., Tischer, D., Chan, F., Koepnick, B., Nguyen, H., Kang, A., Sankaran, B., … Baker, D. (2022). Robust deep learning–based protein sequence design using ProteinMPNN. Science, 378(6615), 49–56. https://doi.org/10.1126/science.add2187 ↩︎
Alcantar, M. A., Paulk, A. M., Moradi, S., Bhar, D., Keller, G. L. J., Sanyal, T., Bai, H., Camdere, G., Han, S. J., Jain, M., Jew, B., Vatansever Inak, S., Langmead, C. J., Tinberg, C. E., Chen, I., & Liu, C. C. (2025). Mapping the evolution of computationally designed protein binders. openRxiv. https://doi.org/10.1101/2025.10.04.680454 ↩︎