Glutamate- and lysine-rich designs are susceptible to expression failure resulting from adenosine-rich sequences
High rates of glutamate and lysine introduction are a staple of structure-based sequence design by ProteinMPNN and related models, regardless of who trains them[1]. Recently, an analysis of the Bits in Bio competition showed that high glutamate/lysine content is predictive of expression failure[2]:
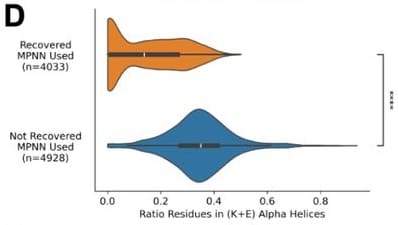
The authors traced this to the fact that both amino acids tend to be encoded by adenosine-rich codons[3], and postulate that high adenosine content led to early termination of translation:
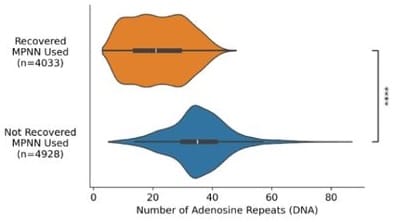
The even-more-recent Proteina-Complexa validation paper[4] showed something similar: when testing designs with phage display, those high in glutamate/lysine content were less likely to be recovered during sequencing:
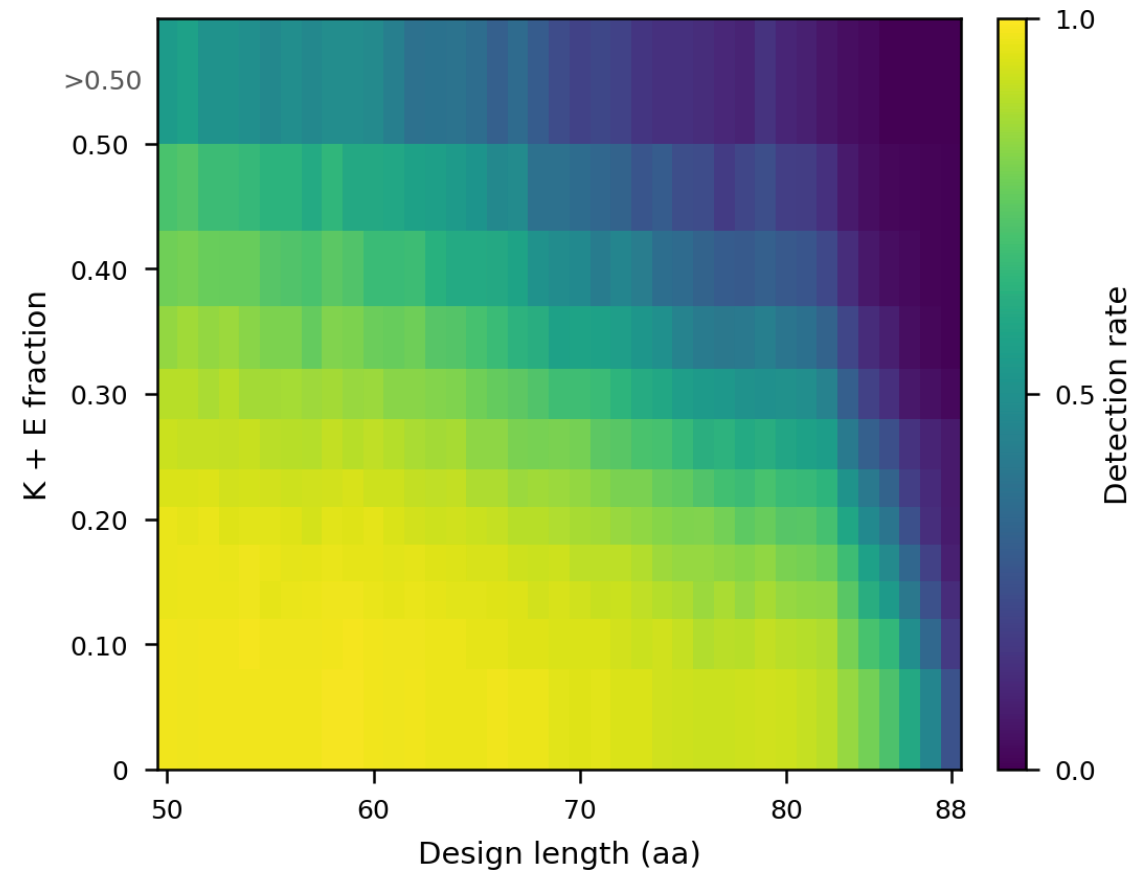
Note that these data do not directly link E/K overinclusion to expression failure. Nevertheless, they do point to the need of additional filters to catch potentially problematic designs.
References
Dauparas, J., Anishchenko, I., Bennett, N., Bai, H., Ragotte, R. J., Milles, L. F., Wicky, B. I. M., Courbet, A., de Haas, R. J., Bethel, N., Leung, P. J. Y., Huddy, T. F., Pellock, S., Tischer, D., Chan, F., Koepnick, B., Nguyen, H., Kang, A., Sankaran, B., … Baker, D. (2022). Robust deep learning–based protein sequence design using ProteinMPNN. Science, 378(6615), 49–56. https://doi.org/10.1126/science.add2187 ↩︎
Stark, H., Faltings, F., Choi, M., Xie, Y., Hur, E., O’Donnell, T., Bushuiev, A., Uçar, T., Passaro, S., Mao, W., Reveiz, M., Bushuiev, R., Pluskal, T., Sivic, J., Kreis, K., Vahdat, A., Ray, S., Goldstein, J. T., Savinov, A., … Jaakkola, T. (2025). BoltzGen: Toward Universal Binder Design. openRxiv. https://doi.org/10.1101/2025.11.20.689494 ↩︎
Kosonocky, C. W., Abel, A. M., Feller, A. L., Cifuentes Rieffer, A. E., Woolley, P. R., Lála, J., Barth, D. R., Gardner, T., Ekker, S. C., Ellington, A. D., Wierson, W. A., & Marcotte, E. M. (2026). Validation and analysis of 12,000 AI-driven CAR-T designs in the Bits to Binders competition. openRxiv. https://doi.org/10.64898/2026.03.03.709355 ↩︎
Didi, K., Zhang, Z., Zhou, G., Reidenbach, D., Cao, Z., Cha, S., Geffner, T., Dallago, C., Tang, J., Bronstein, M. M., Steinegger, M., Kucukbenli, E., Vahdat, A., & Kreis, K. (2026). Scaling atomistic protein binder design with generative pretraining and test-time compute. In The Fourteenth International Conference on Learning Representations. https://openreview.net/forum?id=qmCpJtFZra ↩︎