Language models matter less than search algorithms for difficult protein design tasks
Two papers from the last week show how search algorithms improve the performance of deep learning-based protein design when design restraints are available. The first compared AbLang-2, ESM-2, and other models, finding middling success rates and low variance when used as-is[1]. However, a noticeable performance boost was observed when augmenting AbLang-2 with beam search:
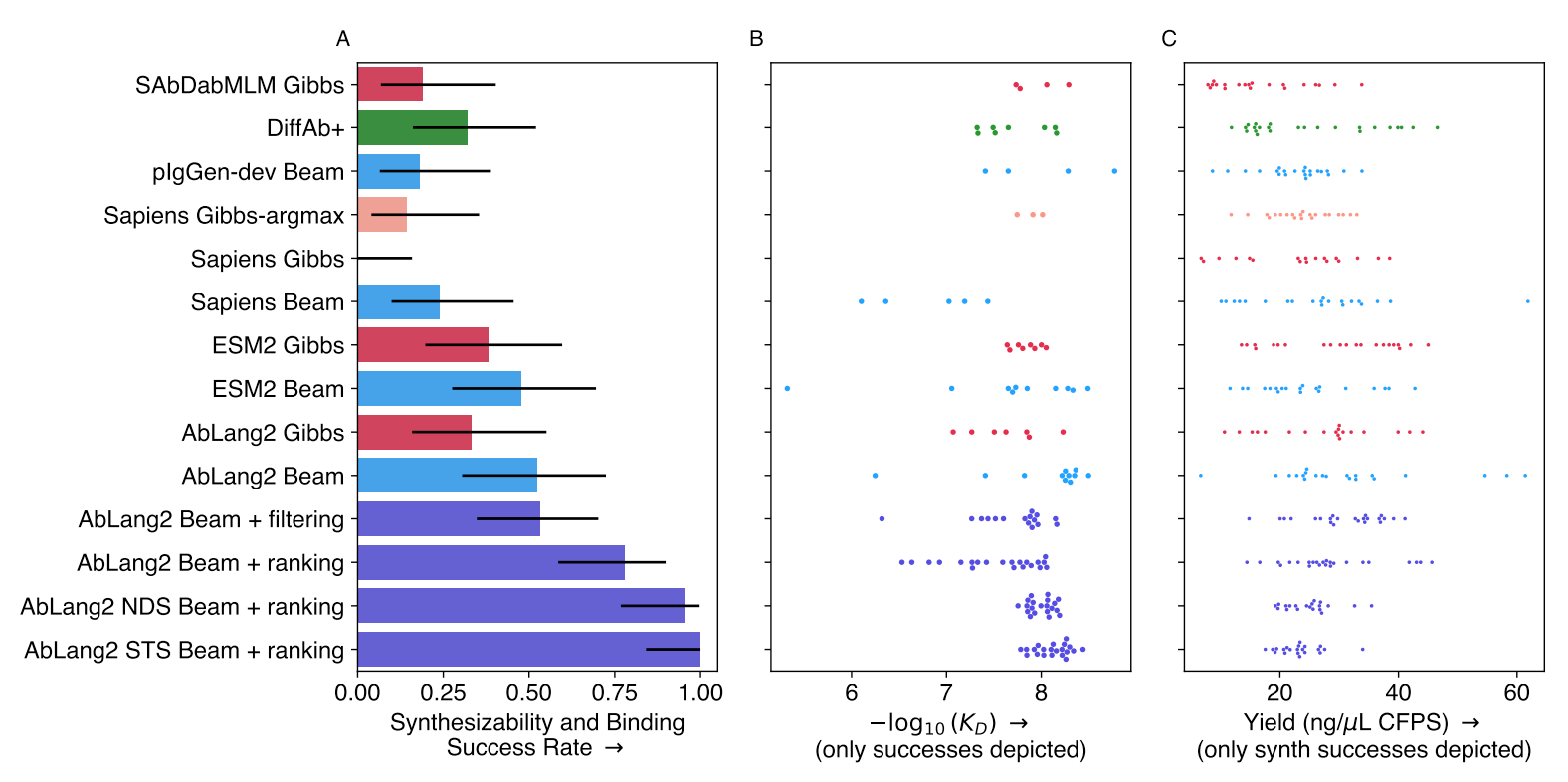
The second, focused on structure-based generative modeling, finds that diverse search algorithms speed up the rate at which diverse solutions are sampled in hard binder design problems[2]:
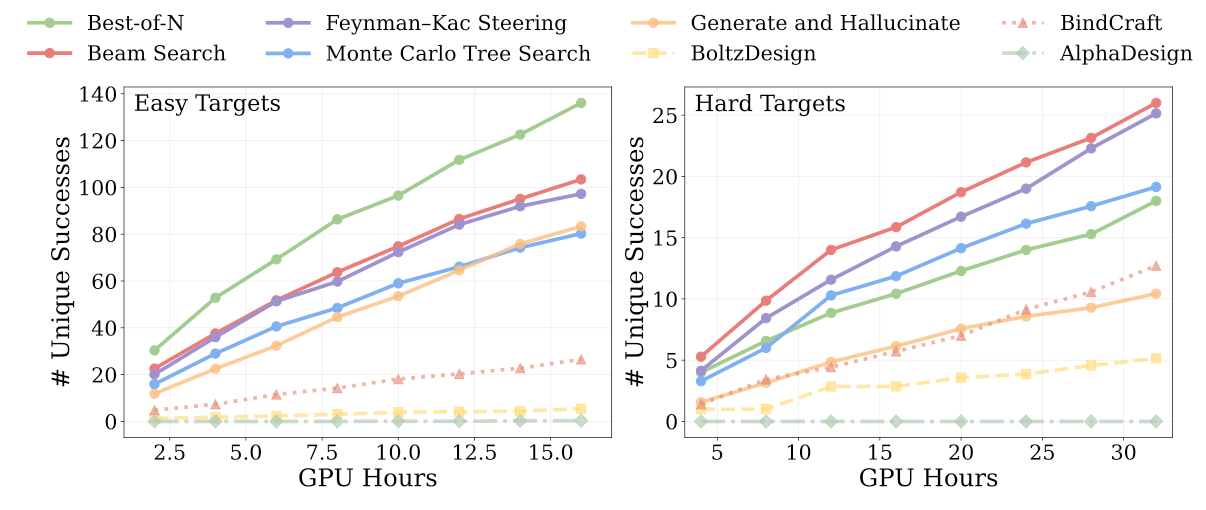
In both cases, the implementations being tested - Gibbs sampling, beam search, Monte Carlo tree search, Feynman-Kac steering - rely on complete rollouts of the design process. A takeaway is that the base model being used probably matters much less than the inclusion (and composition) of potentials, as well as the choice of search algorithms.
References
McCarter, C., Bhattacharya, N., Ober, S. W., & Elliott, H. (2026). How to make the most of your masked language model for protein engineering (Version 1). arXiv. https://doi.org/10.48550/ARXIV.2603.10302 ↩︎
Didi, K., Zhang, Z., Zhou, G., Reidenbach, D., Cao, Z., Cha, S., Geffner, T., Dallago, C., Tang, J., Bronstein, M. M., Steinegger, M., Kucukbenli, E., Vahdat, A., & Kreis, K. (2026). Scaling atomistic protein binder design with generative pretraining and test-time compute. In The Fourteenth International Conference on Learning Representations. https://openreview.net/forum?id=qmCpJtFZra ↩︎