Training inverse folding models with label smoothing improves fitness prediction performance
BERT protein language models and inverse folding models learn to predict masked tokens[1]; coevolutionary patterns and propensities are expected to emerge from training in an unsupervised manner[2]. This works fine when training on sequence data, which is plentiful. However, it is not clear that there are enough experimental structures to safely make this assumption. Label smoothing offers one workaround, where the target distribution being learned for a given residue is not concentrated on the ground truth identity[3]. ProteinMPNN[4] does this, for example, by predicting a distribution of 90% masked (the other 10% is distributed across the other 19 amino acids). I am not aware of any ablation showing benefits of this approach, and the supplement is somewhat cryptic on this matter:

Zhou et al.[5] instead used the BLOSUM62 substitution matrix, a more principled approach than uniform label smoothing. Distributing some density to chemically similar amino acids improved fitness prediction tasks downstream by over 30%:
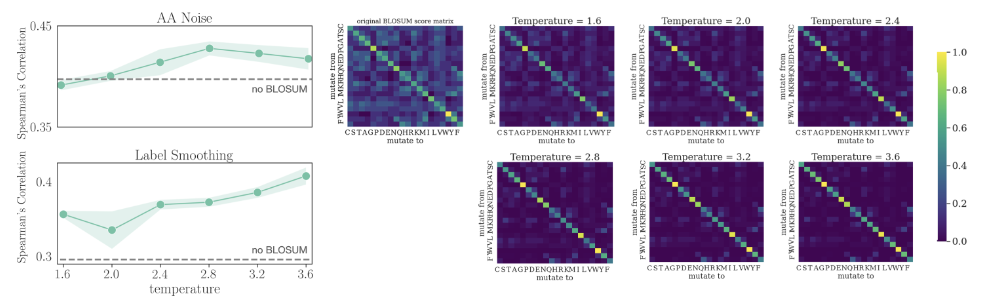
Gong et al.[6] instead use propensities calculated from position-specific scoring matrices, finding that retraining the neural network MutCompute on these propensities led to across-the-board improvements in fitness prediction relative to the original design.
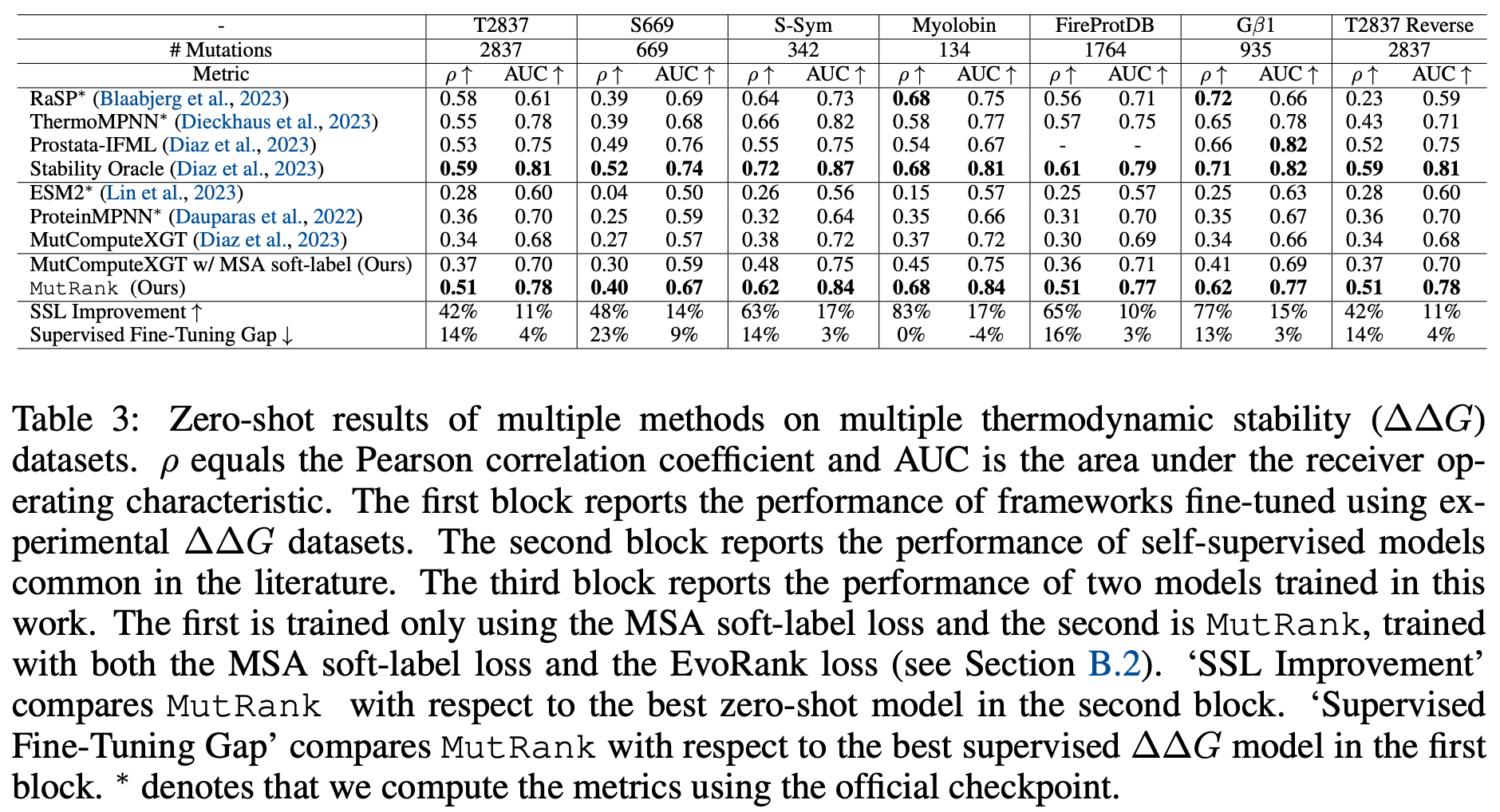
However, the workaround most practitioners have converged upon is to solve the data scarcity issue by augmenting training datasets with synthetic structures. For example, ESM-IF1, perhaps the most widely used inverse folding model[7], has been shown to learn some evolutionary statistics from its exposure to millions of synthetic data points.[8]. Yet a reliance on synthetic data can cause its own issues if experimental structures don't get included during training.
References
Devlin, J., Chang, M.-W., Lee, K., & Toutanova, K. (2018). BERT: Pre-training of Deep Bidirectional Transformers for Language Understanding (Version 2). arXiv. https://doi.org/10.48550/ARXIV.1810.04805 ↩︎
Rives, A., Meier, J., Sercu, T., Goyal, S., Lin, Z., Liu, J., Guo, D., Ott, M., Zitnick, C. L., Ma, J., & Fergus, R. (2021). Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proceedings of the National Academy of Sciences, 118(15). https://doi.org/10.1073/pnas.2016239118 ↩︎
Müller, R., Kornblith, S., & Hinton, G. (2019). When Does Label Smoothing Help? (Version 3). arXiv. https://doi.org/10.48550/ARXIV.1906.02629 ↩︎
Dauparas, J., Anishchenko, I., Bennett, N., Bai, H., Ragotte, R. J., Milles, L. F., Wicky, B. I. M., Courbet, A., de Haas, R. J., Bethel, N., Leung, P. J. Y., Huddy, T. F., Pellock, S., Tischer, D., Chan, F., Koepnick, B., Nguyen, H., Kang, A., Sankaran, B., … Baker, D. (2022). Robust deep learning–based protein sequence design using ProteinMPNN. Science, 378(6615), 49–56. https://doi.org/10.1126/science.add2187 ↩︎
Zhou, B., Zheng, L., Wu, B., Tan, Y., Lv, O., Yi, K., Fan, G., & Hong, L. (2024). Protein Engineering with Lightweight Graph Denoising Neural Networks. Journal of Chemical Information and Modeling, 64(9), 3650–3661. https://doi.org/10.1021/acs.jcim.4c00036 ↩︎
Gong, C., Klivans, A., Loy, J.M., Chen, T., Liu, Q. & Diaz, D.J.. (2024). Evolution-Inspired Loss Functions for Protein Representation Learning. Proceedings of the 41st International Conference on Machine Learning, in Proceedings of Machine Learning Research 235:15893-15906 Available from https://proceedings.mlr.press/v235/gong24e.html. ↩︎
Hsu, C., Verkuil, R., Liu, J., Lin, Z., Hie, B., Sercu, T., Lerer, A., & Rives, A. (2022). Learning inverse folding from millions of predicted structures. openRxiv. https://doi.org/10.1101/2022.04.10.487779 ↩︎
Li, F.-Z., Yang, J., Johnston, K. E., Gürsoy, E., Yue, Y., & Arnold, F. H. (2025). Evaluation of machine learning-assisted directed evolution across diverse combinatorial landscapes. Cell Systems, 16(9), 101387. https://doi.org/10.1016/j.cels.2025.101387 ↩︎