Training protein structure-based neural networks exclusively on predicted protein structures worsens performance on experimental structures due to the training data's idealized local geometry
Predicted protein structures, particularly monomeric structures, have become ubiquitous thanks to the release of the AlphaFold Database[1] and its successors[2]. Yet training structure-based neural networks exclusively on these synthetic structures has now been widely shown to worsen performance on experimental structures.
Hsu et al., who trained the structure-based sequence design neural network ESM-IF1, first noticed that larger models trained only on millions of AlphaFold structures did not generalize to structures in the PDB [3].
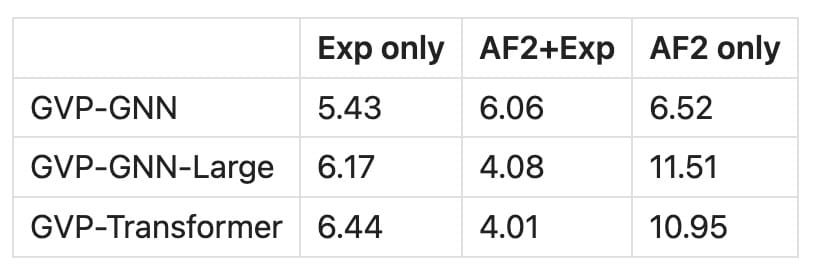
Note: this table is a composite from tables 1 and C.22 from Hsu et al 2022
A more expansive version of this same observation was noted in the SaProt paper[4]. There, several structure-based neural networks were compared, with neural networks trained on synthetic structures failing to generalize to PDB structures:
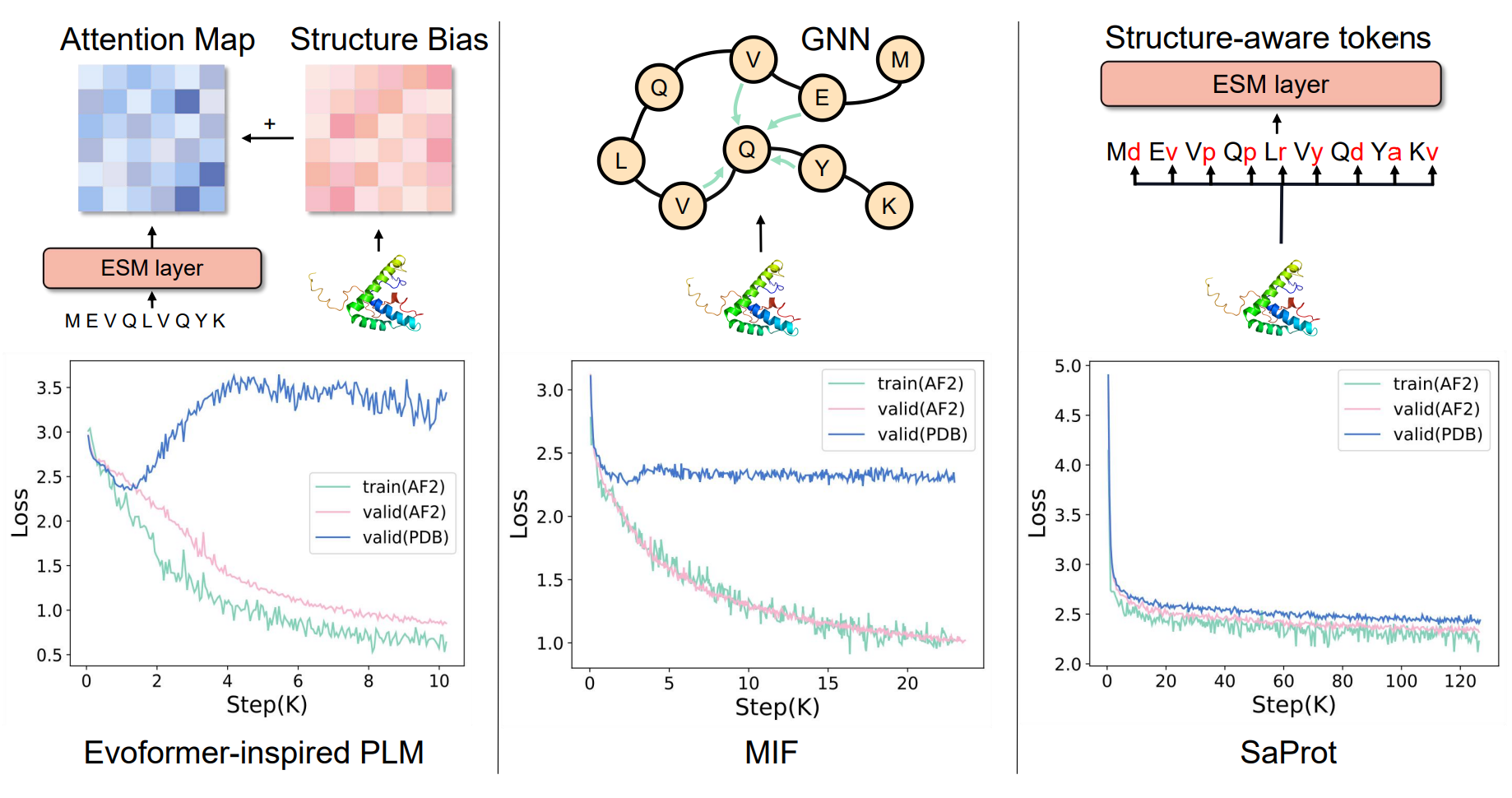
(SaProt, by contrast, trains on tokenized amino acids, rather than all-atom structural data)
The most compelling explanation I've heard was articulated by Tan et al last year[5], which is that these synthetic protein structures only reflect the most common subset of features in the PDB. That includes the most common backbone angles, torsion angles, bond lengths, and so on:
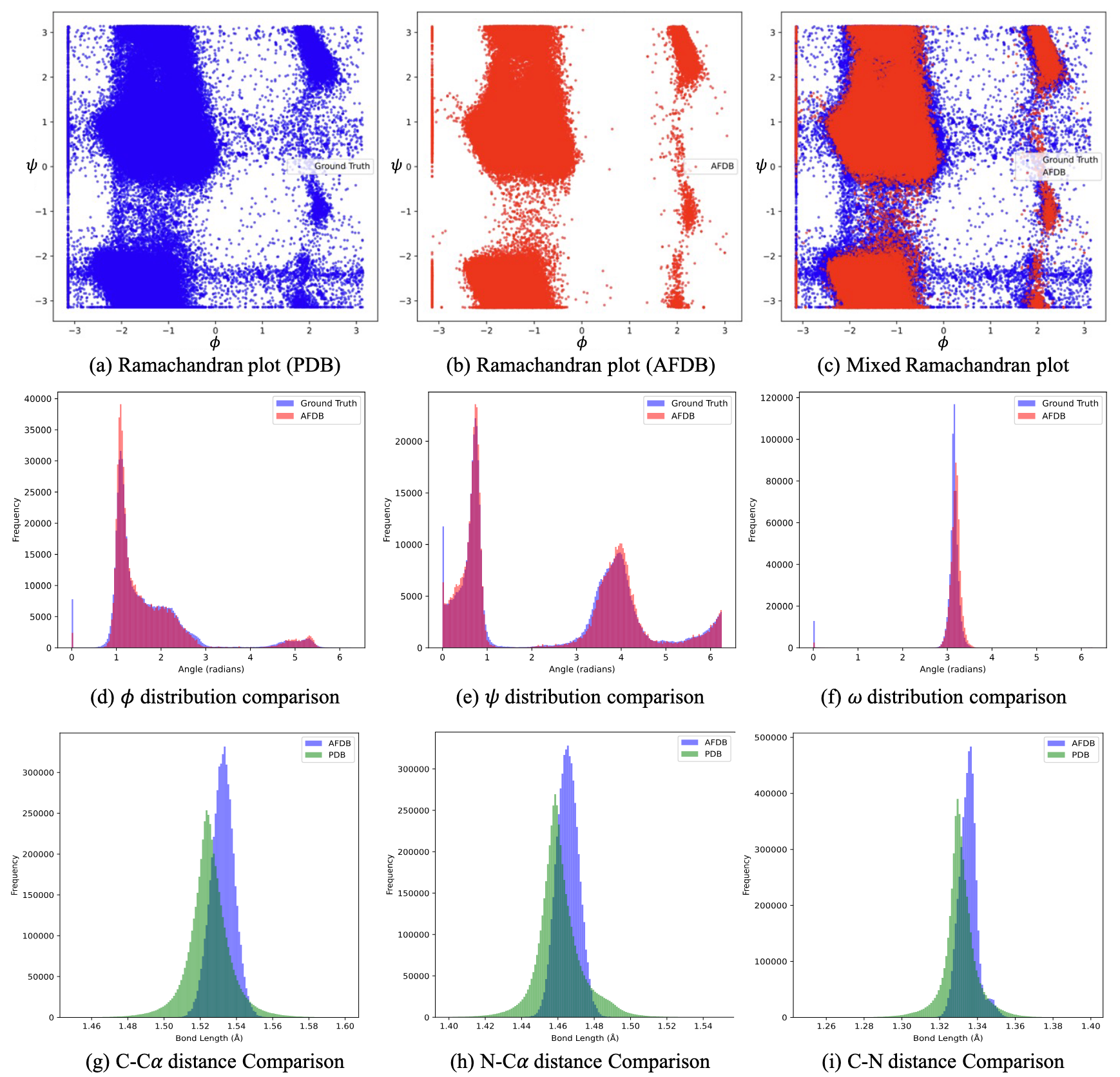
In other words, these synthetic structures are locally perfect, while those deposited in the PDB are not.
References
Tunyasuvunakool, K., Adler, J., Wu, Z., Green, T., Zielinski, M., Žídek, A., Bridgland, A., Cowie, A., Meyer, C., Laydon, A., Velankar, S., Kleywegt, G. J., Bateman, A., Evans, R., Pritzel, A., Figurnov, M., Ronneberger, O., Bates, R., Kohl, S. A. A., … Hassabis, D. (2021). Highly accurate protein structure prediction for the human proteome. Nature, 596(7873), 590–596. https://doi.org/10.1038/s41586-021-03828-1 ↩︎
Lin, Z., Akin, H., Rao, R., Hie, B., Zhu, Z., Lu, W., Smetanin, N., Verkuil, R., Kabeli, O., Shmueli, Y., dos Santos Costa, A., Fazel-Zarandi, M., Sercu, T., Candido, S., & Rives, A. (2023). Evolutionary-scale prediction of atomic-level protein structure with a language model. Science, 379(6637), 1123–1130. https://doi.org/10.1126/science.ade2574 ↩︎
Hsu, C., Verkuil, R., Liu, J., Lin, Z., Hie, B., Sercu, T., Lerer, A., & Rives, A. (2022). Learning inverse folding from millions of predicted structures. openRxiv. https://doi.org/10.1101/2022.04.10.487779 ↩︎
Su, J., Han, C., Zhou, Y., Shan, J., Zhou, X., & Yuan, F. (2023). SaProt: Protein Language Modeling with Structure-aware Vocabulary. openRxiv. https://doi.org/10.1101/2023.10.01.560349 ↩︎
Tan, C., Cao, Z., Gao, Z., Li, S., Huang, Y., & Li, S. Z. (2025). AlphaFold Database Debiasing for Robust Inverse Folding (Version 1). arXiv. https://doi.org/10.48550/ARXIV.2506.08365 ↩︎